1. INTRODUCTION
The increasing human population and climate change are causing mounting pressure on food security which is dependent on the agricultural sector, particularly food crops. The world’s human population is estimated to reach 8.6 billion by the year 2030 [1] and this will lead to a widening gap between food demand and production. Adding to this problem is the anthropogenic activities that have led to climate change resulting in biotic and abiotic stresses in crop plants, which eventually will lead to reduced crop production [1]. Thus, climate change will further widen the food demand and production gap.
Beneficial microorganisms, which can be obtained from within plants or from their rhizospheres, can be used to improve plant growth and crop yield without harming the environment. These microorganisms can directly or indirectly promote plant growth by increasing the availability of nutrients in the soil, fixing nitrogen and making it available to plants, modulating plant growth through the production of growth regulators, and producing compounds that are antagonistic to other pathogenic microorganisms. For many years, rhizosphere bacteria have been used as biofertilizers to promote plant growth and increase crop yield. Plant growth promotion mechanisms mediated by rhizosphere bacteria have been thoroughly studied and elucidated [2], leading to the development of several commercial biofertilizers [3]. Among the rhizosphere bacteria, the Actinomycetota have recently gained importance as plant growth promoters. The phylum Actinomycetota [4] (previously Actinobacteria) are Gram-positive or Gram-variable bacteria that have a rigid cell wall. They are phenotypically diverse ranging from cocci to highly differentiated mycelia with DNA G + C content ranging from 50 mol% to 70 mol% [5]. The phylum has long been known for its ability to produce myriads of metabolites with antimicrobial properties and many of these compounds have reached drug stores today as antibiotics [6]. Actinomycetota isolated from the rhizospheres of various plants have been found to have traits that promote plant growth and biocontrol capability against different phytopathogens [7-11].
Medicinal plants have played important role in the life and health of people for centuries. Some of the bioactive compounds that give medicinal plants their therapeutic benefits are produced by the associated microorganisms or arise from their interactions [12]. However, research into the interactions of medicinal plants and their associated microorganisms is still in its infancy. Microorganisms associated with medicinal plants are also found to have plant growth-promoting (PGP) and biocontrol properties [12]. The previous studies have also shown the PGP potential of Actinomycetota from medicinal plant rhizospheres [13-15].
Two plants, Rubus ellipticus Sm. and Ageratina riparia (Regel) King and Rob., were chosen for this study because they have been used for medicinal purposes by the indigenous people of this region, which is located in the state of Meghalaya, India. R. ellipticus is a medicinal plant in the family Rosaceae. Its various parts are used for the treatment of diabetes, jaundice, hepatitis, stomach ache, fever, cough, sore throat, bone fracture, tumor, ulcer, bacterial infection, abdominal pain, colic pain, epilepsy, urinary tract infection, dysentery, and coronary heart diseases. The plant is also used as a laxative, diuretic, astringent, antifertility, analgesic, antimicrobial, and kidney tonic [16]. A. riparia is an invasive weed in the family Asteraceae and native to Mexico. Its parts are medicinally used to reduce hypertension, blood sugar levels, and blood pressure, and it also has diuretic effects and possibly anti-infective properties against urinary-tract infections [17,18]. Both plants have not been investigated concerning their associated rhizosphere Actinomycetota. This investigation will, therefore, provide some insights into the occurrence of Actinomycetota from the rhizosphere of these two medicinal plants and it would also shed some light on their PGP potential on other food crops such as tomato, which is an important horticultural food crop worldwide.
The purpose of this study is, therefore, to isolate Actinomycetota from the rhizosphere of two medicinal plants, R. ellipticus and A. riparia, and to evaluate their antimicrobial activity against bacterial and fungal strains as well as their ability to promote plant growth both in vitro and in vivo on tomato plants.
2. MATERIALS AND METHODS
2.1. Sample Collection
R. ellipticus plant samples were collected from a forest in Mawklot village, Upper Shillong, Meghalaya (N25°33’07.9”, E91°49’49.2”). A. riparia plant samples were collected from St. Edmund’s College campus, Shillong, Meghalaya (N25°34’00.3”, E91°53’48.2”). Entire plant along with the soil attached to the roots was carefully uprooted, stored in sterilized polythene bags, and brought to the laboratory. The loosely attached soil was shaken off and discarded. The more tightly bound soil was shaken more vigorously, aseptically collected, and treated as rhizosphere soil, and stored at 4 ± 1°C until use.
2.2. Determination of Soil pH and Organic Matter
Soil pH was measured by suspending 10 g fresh unsieved soil in distilled water in a ratio of 1:5 (w/v), stirred for 15 min, followed by another 15 min of settling down of soil particles, after which pH reading was taken. Soil organic matter was estimated using the method of Walkley and Black [19].
2.3. Pre-treatment and Isolation of Actinomycetota
Soil samples were air-dried at room temperature for 1 week, half of which were subsequently dry heated at 120 ± 2°C for 1 h. 1 g of both soil samples was suspended in 9 ml sterilized distilled water and serially diluted up to 103 times. Three different selective media, namely, Actinomycetes isolation agar, starch casein agar, and Streptomyces agar, were inoculated with 100 ml of diluted soil suspension. These media were supplemented with cycloheximide (50 mg ml-1) and rifampicin (20 mg ml-1) to prevent the growth of contaminating fungi and bacteria. Inoculated media were incubated at 28 ± 2°C for a duration of 3–4 weeks and putative Actinomyces total colonies were selected and maintained in Bennett’s agar medium.
2.4. Morphological Characterization
The selected colonies were inoculated on coverslips obliquely inserted on ISP3 media by following the method of Cross [20], incubated at 28°C for 7 days, and observed under a microscope (Olympus CX21i) at ×400. Morphological characterization of the cultures was based on aerial and substrate mycelia and spore chains.
2.5. Molecular Characterization
Genomic DNA was isolated from the cultures by an enzymatic method of Chen et al. [21] and subsequently used for amplification of 16S rRNA gene. Five primer pairs were used depending on their success of amplification, namely, 27F-A3R, 27F-1492R, 27F-Sm5R [22], 27F-1392R [23], and 27F-1525R [24]. The 25 ml PCR mix consisted of PCR buffer with 1.5 mM MgCl2, each dNTP of 0.2 mM, each primer of 0.5 mM, Taq DNA polymerase of 0.625 U, and 5 ml of genomic DNA. Reactions were performed at an initial denaturation temperature of 95°C for 5 min, 35 cycles of denaturation at 95°C for 45 s, annealing temperatures at 51°C, 49°C, 49°C, 50°C, and 48°C for 45 s for the above respective primer pairs, extension temperature at 72°C for 90 s, and final extension of 72°C for 7 min. PCR products were verified by agarose gel electrophoresis (1.2% w/v agarose added with 1 ml/10 ml v/v LabSafe™ Nucleic Acid Stain, G-Biosciences), followed by purification and sequencing at Eurofins Genomics, India. The obtained sequences were compared with sequences from NCBI database for genus identification.
2.6. Nucleotide Accession Numbers
The partial 16S rRNA gene sequences were deposited in GenBank against which the following accession numbers were assigned: MH424581-MH424592 and MH425315-MH425321.
2.7. Antimicrobial Activities
The culture filtrates were evaluated for antibacterial activity against Escherichia coli MTCC 1669, Pseudomonas aeruginosa MTCC 4673, Staphylococcus aureus MTCC 9886, Bacillus subtilis MTCC 1305, and anticandidal activity against Candida albicans MTCC 7253 by following the agar well diffusion method described by Syiemiong and Jha [25], except that the incubation was carried out under stationary condition at 30 ± 2°C for 30 days. The cultures were also evaluated for antagonistic activity by dual culture against phytopathogenic bacterial and fungal strains, namely, Xanthomonas campestris pv. campestris Pammel (ITCC BH0004), X. oryzae pv. oryzae Ishiyama (ITCC BB0013), Ralstonia solanacearum Smith (ITCC BI0001), Fusarium oxysporum f. sp. zingiberi Trujillo (ITCC 2698), and Aspergillus niger (MTCC 4325). Dual culture assay against bacterial phytopathogens was performed according to Vijayakumar et al. [26] and against fungal phytopathogens, according to Khamna et al. [27]. MTCC strains were obtained from IMTECH, Chandigarh, India, and ITCC strains were obtained from IARI, New Delhi, India.
2.8. Detection of Biosynthetic Gene Clusters (BGCs)
Detection of BGCs, namely, Type I polyketide synthase (PKS-I), type II polyketide synthase (PKS-II), and non-ribosomal peptide synthetase was carried out according to the method described by Syiemiong and Jha [25].
2.9. In vitro PGP Activities
Indole-3-acetic acid (IAA) production, phosphate solubilization, siderophore, and diazotrophic activities were evaluated according to the methods described by Syiemiong and Jha [28].
2.10. Tomato Seed Germination Assay and Seedling Growth
Tomato seeds (Rocky, Syngenta) were surface sterilized with 70% ethanol for 5 min, followed by 2% NaOCl for 3 min, and rinsed 5 times in sterilized distilled water. Ten surface-sterilized seeds were soaked in five selected 2-week Actinomycete suspension cultures (stationary condition; 105 CFU ml-1) for 2 h and subsequently transferred into sterilized Petri dishes containing sterilized moistened filter paper and incubated at 25 ± 2°C until germination. Control seeds were surface sterilized as above, but soaked in uninoculated broth and incubated in the same way. The Petri dishes were regularly moistened with sterilized distilled water. Germination percentage was measured as number of seeds germinated out of the total number of seeds ×100. The length of the newly germinated seedlings was also measured 4 days post-germination. The vigor index of tomato seedlings was measured according to Tamreihao et al. [29].
2.11. PGP Assay on Tomato Plant under Different Soil Conditions
To study the influence of resident microorganisms on the PGP activity of the selected isolates, tomato plants were grown on sterilized and unsterilized soil conditions. To study the influence of organic matter, the tomato plants were grown on soil amended with and without organic manure. To perform the above investigations, the assay was conducted in four different soil conditions, namely, (1) sterilized soil, (2) unsterilized soil, (3) sterilized soil amended with 30% (w/w) sterilized organic manure, and (4) unsterilized soil amended with 30% (w/w) unsterilized organic manure. Commercially produced organic manure was used with the following composition: 30–40% moisture, 90% of the material pass through 4 mm sieve, bulk density (g/cm3) <1.0, total viable count (N, P, K, Zn bacteria or N, P bacteria or N, K bacteria) of 5.0 × 106, total organic carbon of 14%, total nitrogen (N) of 0.8%, total phosphorus (P2O5) of 0.5%, total potassium (K2O) of 0.8%, total NPK of 3%, conductivity (dsm-1) <4, pH 6.5–8.0, C: N ratio <16, and heavy metal content (mg/kg) As2O3 10, Cd 5, Cr 50, Cu 300, Hg 0.15, Ni 50, Pb 100, and Zn 1000. Under sterilized soil condition, tomato seeds were surface sterilized as already described, followed by sowing on germination trays filled with sterilized peat. Germinated seedlings were transplanted into plastic cups containing soil as in (1) or (3) above. Under unsterilized soil condition, the germinated seedlings were transplanted into cups containing soil as in (2) and (4) above. Immediately after transplantation, the seedlings were soil drenched by the selected 2-week Actinomycete suspension cultures (105 cfu ml-1) around the base of the stem. Control plants were drenched with uninoculated broth. The second drenching was done 1 week later and the plants were finally uprooted 2 weeks later for the measurement of plant fresh weight. All treatments were conducted in five replicates. All the plants were kept in a completely randomized fashion under a net house (50% shade) with daily mid-day temperatures ranging from 21.23 ± 0.77°C to 27.90 ± 0.91°C and relative humidity from 54.33 ± 4.67% to 99.00 ± 0.00%.
2.12. Statistical Analysis
Multiple comparisons between sample means and control means were performed by ANOVA and Tukey’s HSD test at P < 0.05. Statistical analysis of the data was done using SPSS.
3. RESULTS AND DISCUSSION
3.1. Morphological and Molecular Characterization
Fifty-one putative colonies from the rhizosphere of R. ellipticus and 42 colonies from that of A. riparia were selected, out of which 16 from R. ellipticus and 11 from A. riparia were verified as Actinomycetota according to the micromorphological characteristics. Twelve isolates from the rhizosphere of R. ellipticus and seven from that of A. riparia could be identified using 16S rRNA gene sequence. The remaining isolates were mostly identified only on the basis of micromorphological characteristics due to their poor genomic DNA quality. Overall, Streptomyces was the dominant genus from both the rhizospheres, albeit Amycolatopsis was also prominent from the rhizosphere of R. ellipticus [Figure 1]. The isolates, their GenBank accession numbers, and their closest neighbors from the NCBI database are given in Table 1. The high frequency of isolation of Streptomyces, as shown in this study, has also been reported in other rhizospheres of medicinal and crop plants from India [30-32] and other parts of the world [27,33,34]. Streptomyces is found to be able to easily adapt to various nutritional conditions and availability [35] due to its metabolic versatility, and this could be one of the reasons for its fast growth and colonization on artificial media during isolation. Apart from Streptomyces, Amycolatopsis has also been reported as a frequent genus isolated from soils and rhizospheres of numerous plant species [33,36-39].
 | Figure 1: Actinomycete genera from rhizospheres of A. riparia and R. ellipticus. [Click here to view] |
Table 1: Identified actinomycete isolates from rhizospheres of A. riparia and R. ellipticus.
| Isolate | GenBank accession No.a | Closest type strain from NCBI databaseb | Percent identity | Query cover |
|---|---|---|---|---|
| AR-10 | MH425315 | Streptomyces capoamus NBRC 13411 (NR_112394) | 99.27% | 99% |
| AR-11 | - | Amycolatopsis sp. | - | - |
| AR-14 | - | Streptomyces sp. | - | - |
| AR-23 | MH425316 | Streptomyces umbrinus NRRL B-2572 (NR_115792) | 99.54% | 100% |
| AR-24 | MH425317 | Streptomyces bottropensis ATCC 25435 (NR_115571) | 97.57% | 100% |
| AR-27 | MH425318 | Amycolatopsis umgeniensis UM16 (NR_115688) | 91.50% | 100% |
| AR-28 | MH425319 | Streptomyces umbrinus NRRL B-2572 (NR_115792) | 94.19% | 90% |
| AR-30 | MH425320 | Streptomyces umbrinus NRRL B-2572 (NR_115792) | 98.71% | 99% |
| AR-34 | - | Streptomyces sp. | - | - |
| AR-37 | MH425321 | Streptomyces umbrinus NRRL B-2572 (NR_115792) | 99.64% | 100% |
| RE-05 | MH424581 | Amycolatopsis vastitatis H5 (NR_164904) | 99.84% | 100% |
| RE-13 | MH424582 | Amycolatopsis vastitatis H5 (NR_164904) | 100% | 100% |
| RE-17 | MH424583 | Streptomyces cacaoi subsp. asoensis NBRC 13813 (NR_112416) | 99.47% | 100% |
| RE-19 | MH424584 | Streptomyces yatensis NBRC 101000 (NR_041427) | 98.62% | 100% |
| RE-20 | MH424585 | Streptomyces kronopolitis NEAU-ML8 (NR_153682) | 99.29% | 100% |
| RE-22 | MH424586 | Amycolatopsis roodepoortensis M29 (NR_134695) | 98.80% | 100% |
| RE-27 | MH424587 | Streptomyces melanosporofaciens NRRL B-12234 (NR_114819) | 99.80% | 100% |
| RE-29 | MH424588 | Streptomyces cacaoi subsp. asoensis NBRC 13813 (NR_112416) | 99.69% | 100% |
| RE-31 | MH424589 | Streptomyces cacaoi subsp. asoensis NBRC 13813 (NR_112416) | 100% | 100% |
| RE-32 | MH424590 | Amycolatopsis vastitatis H5 (NR_164904) | 99.82% | 100% |
| RE-34 | MH424591 | Streptomyces prunicolor NBRC 13075 (NR_112359) | 98.94% | 99% |
| RE-39 | MH424592 | Promicromonospora callitridis CAP94 (NR_158047) | 99.35% | 100% |
| RE-42 | - | Amycolatopsis sp. | - | - |
| RE-44 | - | Amycolatopsis sp. | - | - |
(a): The isolates with GenBank accession number were identified using 16S rRNA gene sequence, and those without GenBank accession number were putatively identified by micromorphological characteristics only. (b): The type strains from NCBI database were given as species, followed by strain name and GenBank accession number in parenthesis.
3.2. Antimicrobial Activities
Agar well diffusion assay of the culture filtrates showed nine isolates from the rhizosphere of R. ellipticus with antimicrobial potential against the bacterial and fungal strains, while there were almost no isolates from the rhizosphere of A. riparia that showed antimicrobial activity. Dual culture assays against bacterial and fungal phytopathogens also showed more numbers of isolates from the rhizosphere of R. ellipticus with antagonistic activity. However, looking at their genetic potential for antimicrobial properties, the distribution of BGCs related to the synthesis of antimicrobial compounds was more or less the same from the isolates of the two rhizospheres. Despite the similarity in BGC profile, the phenotypic difference in expressing antimicrobial activity between the isolates of the two rhizospheres could be due to the difference in organic matter of the soils where the plants were growing. It was previously found that composts can shift the microbial community structure of the rhizosphere, thereby increasing the incidence of bacteria antagonistic against soil-borne root pathogens [40]. In this study, the soils from the two sampling sites had significantly different levels of organic matter. The forest soil from where R. ellipticus plants were collected had more organic matter (158.31 ± 14.20 mg g-1 soil) than that of the college campus soil from where A. riparia plants were collected (96.06 ± 1.07 mg g-1 soil). The higher organic matter of the forest soil could have changed its microbiome structure, thereby increasing the incidence of isolation of antagonistic Actinomycetota. There was also a difference in pH of the soils from the two sites, with the forest soil being acidic (pH 4.52 ± 0.03) and the college campus soil being alkaline (8.07 ± 0.09). The reason for the better recovery of antimicrobial-producing strains from acidic soil in this work is not clear, but the previous studies have also reported on antimicrobial-producing Actinomycetota from acidic soils [41,42]. Among the antagonistic isolates from this study, they were mostly of the genus Amycolatopsis and a few from Streptomyces. Streptomyces is already a well-known antibiotic-producing genus of Actinomycetota. The genus Amycolatopsis has also been reported to be a potent antibiotic producer [33,43] and several bioactive compounds have been characterized from it [44-46]. The isolates which showed antagonistic potential in vitro were, however, not tested in vivo for biocontrol activity against disease-causing phytopathogens, and this evaluation needs to be looked into.
3.3. In vitro PGP Activities
In vitro PGP assays of the isolates in terms of IAA production, phosphate solubilization, siderophore, and diazotrophic activities, however, showed that the isolates from the rhizosphere of A. riparia showed better PGP properties as compared to those from the rhizosphere of R. ellipticus. The difference in PGP activities among the isolates from these two rhizospheres could again be due to reasons with respect to soil pH and organic matter. The alkaline nature of the soil from the college campus as already mentioned, combined with its scanty vegetation, could have been responsible for its low organic matter and hence its low nutrient availability [47]. This could have probably assembled PGP microorganisms to enhance nutrient availability. Other workers have also reported on the isolation of plant growth promoting rhizobacteria (PGPR) strains from nutrient-limited soils such as from desert and semi-desert conditions [48,49]. The stark difference in PGP properties among the isolates from the two rhizospheres in this study could also be due to the plants themselves assembling specific microbial communities with specific PGP activities due to the differences in the root exudates and chemical signals secreted by them [50]. Almost all the isolates with PGP property belonged to the genus Streptomyces. PGP Streptomyces strains have also been routinely isolated from rhizospheres and endospheres of numerous plants by previous workers. Al-Dhabi et al. isolated Streptomyces strains from the rhizosphere of tomato plants which could produce IAA, solubilize phosphate, and produce siderophores [51]. Barman and Dkhar isolated numerous PGP Streptomyces strains from the endosphere of several medicinal plants from Meghalaya [52,53].
3.4. Tomato Seed Germination Assay and Seedling Growth
Based on overall PGP activities assayed in vitro, broth cultures of four isolates from the rhizosphere of A. riparia (Amycolatopsis AR-11, Streptomyces AR-23, Streptomyces AR-24, and Streptomyces AR-37) and one from the rhizosphere of R. ellipticus (Streptomyces RE-29) were tested in vitro for their ability to increase seed germination and to enhance seedling growth in tomato. All the five isolates significantly increased seed germination and seedling growth, and hence increased the vigor of the tomato seedlings [Figure 2]. Three isolates, namely, Streptomyces AR-23, Streptomyces AR-37, and Streptomyces RE-29 showed maximum vigor which were not significantly different from each another. A tabulated comparison of the PGP activity of the tested isolates on seed germination percentage, seedling length, and vigor index of tomato is given in Table 2. Other workers have also reported on Streptomyces strains showing enhancing effects on seedling growth of plants. An endophytic Streptomyces strain BPSAC147 isolated from Rhynchotechum ellipticum in Mizoram was found to significantly enhance seedling growth of tomato in vitro [54]. Barman and Dkhar also reported endophytic Streptomyces strains isolated from medicinal plants in Meghalaya that could increase root and shoot length in rice plants [53]. Dochhil et al. reported on two endophytic Streptomyces strains isolated from Centella asiatica in Meghalaya that significantly enhanced seed germination and seedling growth in Phaseolus vulgaris [55].
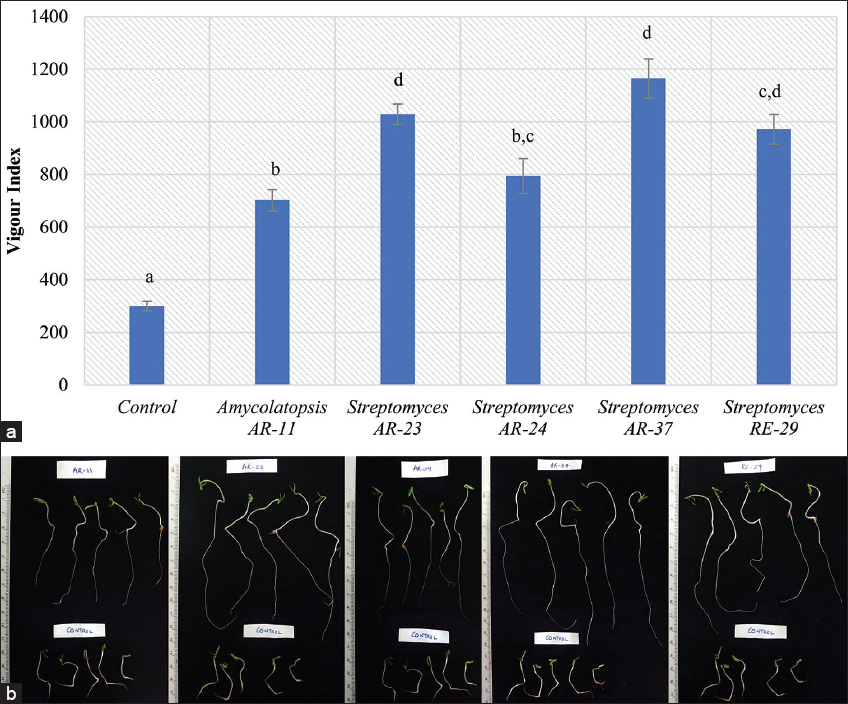 | Figure 2: (a) Vigor indices of tomato seedlings after treatment with broth cultures of the selected isolates; vertical bars = standard error; means with different letters above vertical bars are significantly different from each other at P < 0.05. B 4-day-old tomato seedlings after treatment with broth cultures of the selected PGP isolates. (b). 4-day-old tomato seedlings after treatment with broth cultures of the selected PGP isolates. [Click here to view] |
Table 2: Effect of the selected isolates on seed germination, seedling length, and vigor index of tomato.
| Treatment | Genus | Seed germination (%) | Seedling length (cm) | Vigor index |
|---|---|---|---|---|
| Control | - | 71.43 | 4.20±0.26a | 300.00±18.48a |
| AR-11 | Amycolatopsis | 87.50 | 8.03±0.46b | 702.50±40.54b |
| AR-23 | Streptomyces | 93.33 | 11.02±0.42d | 1028.67±38.84d |
| AR-24 | Streptomyces | 92.86 | 8.55±0.71b,c | 794.29±65.49b,c |
| AR-37 | Streptomyces | 100.00 | 11.65±0.75d | 1164.67±74.65d |
| RE-29 | Streptomyces | 93.33 | 10.41±0.61c,d | 972.00±56.73c,d |
Different superscript letters among different treatments under seedling length and vigor index indicate significant differences at P<0.05. Values under seedling length and vigor index are given as mean±standard error. Fifteen replicates have been taken for each observation.
3.5. PGP Assay on Tomato Plant under Different Soil Conditions
To test the ability of the five selected isolates to promote plant growth in vivo, tomato plants were used as test plants. The tomato plants were grown in four different garden soil conditions as already described, namely, sterilized soil, unsterilized soil, sterilized soil and manure, and unsterilized soil and manure. The results of pH and organic matter analysis of the above soil conditions are given in Table 3. All the four soil conditions fall under the optimum pH range required for normal plant growth, which is usually from 6.1 to 7.8, a range associated with optimum availability of soil nutrients [47]. However, the organic matter was significantly more in the soils amended with manure. There were no significant differences in organic matter between sterilized and unsterilized soils.
Table 3: pH and organic matter of garden soil under different soil conditions.
| Soil condition | Soil pH | Organic matter (mg g-1 soil) |
|---|---|---|
| Sterilized soil | 6.61±0.02b | 56.35±0.00a |
| Unsterilized soil | 7.02±0.04d | 61.71±5.37a |
| Sterilized soil and manure | 6.46±0.02a | 131.48±10.73b,c |
| Unsterilized soil and manure | 6.76±0.02c | 147.58±10.73c |
Different superscript letters among different soil conditions under pH and organic matter indicate significant differences at P<0.05. Values are given as mean±standard error. Three replicates were taken for each observation.
Without the amendment of organic manure and under sterilized soil condition, none of the tested isolates could significantly enhance tomato plant growth. However, under unsterilized soil condition, three of the tested isolates, namely, AR-11, AR-23, and RE-29 could significantly enhance plant growth in terms of plant fresh weight [Table 4]. With the amendment of organic manure, one isolate AR-37 could significantly enhance tomato plant growth under sterilized soil condition and two more isolates AR-11 and AR-23 in addition to AR-37 could also significantly enhance plant growth under unsterilized soil condition [Table 5]. These results indicated a positive influence of resident soil microorganisms in enhancing PGP activity of the tested isolates under soils of both low and high organic matter. Hassani et al. [56] also reviewed on the existence of “helper” microbes which positively influenced or assisted other PGP microbes in promoting plant growth. Although the helper effect of the resident soil microorganisms on the tested PGP isolates were found to be significant under soils not amended with organic manure [Table 4], the helper effect was, however, not significant on soils amended with organic manure [Table 5]. The reason could be that the presence of abundant organic matter in the soil surpassed the requirement of the resident soil microorganisms in helping the PGP isolates to promote plant growth, and their requirement to help the PGP isolates becomes essential only under low nutrient conditions. Arif et al. [57] also showed that PGPR combined with nitrogen-enriched compost could significantly improve productivity and seed quality in sunflowers.
Table 4: Plant growth promotion of tomato plants by selected isolates under sterilized and unsterilized soil conditions without the amendment of organic manure.
| Treatment | Soil condition | Plant fresh weight (g) |
|---|---|---|
| Control | Sterilized | 0.08±0.01a,b |
| Unsterilized | 0.15±0.02a,b,c | |
| Amycolatopsis AR-11 | Sterilized | 0.10±0.01a,b |
| Unsterilized | 0.27±0.04e | |
| Streptomyces AR-23 | Sterilized | 0.11±0.01a,b,c |
| Unsterilized | 0.24±0.01d,e | |
| Streptomyces AR-24 | Sterilized | 0.07±0.01a |
| Unsterilized | 0.16±0.02b,c,d | |
| Streptomyces AR-37 | Sterilized | 0.08±0.01a,b |
| Unsterilized | 0.19±0.01c,d,e | |
| Streptomyces RE-29 | Sterilized | 0.09±0.01a,b |
| Unsterilized | 0.24±0.03d,e |
Different superscript letters after plant fresh weight indicate significant differences at P<0.05. Values are given as mean±standard error. Five replicates were taken for each observation.
Table 5: Plant growth promotion of tomato plants by selected isolates under sterilized and unsterilized soil conditions with the amendment of organic manure.
| Treatment | Soil condition | Plant fresh weight (g) |
|---|---|---|
| Control | Sterilized | 0.28±0.05a,b |
| Unsterilized | 0.24±0.04a | |
| Amycolatopsis AR-11 | Sterilized | 0.63±0.09a,b,c,d,e |
| Unsterilized | 0.74±0.06b, c, d, e | |
| Streptomyces AR-23 | Sterilized | 0.75±0.14b,c,d,e |
| Unsterilized | 1.05±0.10e | |
| Streptomyces AR-24 | Sterilized | 0.48±0.03a,b,c |
| Unsterilized | 0.56±0.08a,b,c,d | |
| Streptomyces AR-37 | Sterilized | 0.85±0.19c,d,e |
| Unsterilized | 0.99±0.11d,e | |
| Streptomyces RE-29 | Sterilized | 0.63±0.10a,b,c,d,e |
| Unsterilized | 0.57±0.08a,b,c,d |
Different superscript letters after plant fresh weight indicate significant differences at P<0.05. Values are given as mean±standard error. Five replicates were taken for each observation.
The influence of different treatments and soil conditions on PGP activity of tomato plants is shown in [Figure 3a and b], and 25-day-old tomato plants treated with Streptomyces AR-23 are shown in [Figure 3c]. Although some of the tested PGP isolates showed promising results on tomato plant growth, these results are still preliminary to be conclusive, and hence, further intensive investigations are needed under field conditions to assess their effects on tomato crop productivity on a larger scale.
 | Figure 3: (a-b) Comparison of growth of tomato plant under different treatments and soil conditions; vertical bars = standard error. (c) A 25-day-old tomato plant treated with Streptomyces AR-23 under different soil conditions [Click here to view] |
4. CONCLUSION
Streptomyces was the most abundant genus isolated from the rhizospheres of the two medicinal plants R. ellipticus and A. riparia. The rhizosphere of R. ellipticus was found to harbor more antagonistic Actinomycetota (mostly Amycolatopsis) while that of A. riparia harbored more PGP Actinomycetota (mainly Streptomyces). Soil pH and organic matter showed indications of influencing antimicrobial and PGP properties of the rhizosphere-dwelling Actinomycetota. The five selected isolates significantly increased the vigor of tomato seedlings in vitro and could also improve tomato plant growth in vivo under net house conditions under abundant organic matter. The resident soil microorganisms could also help the selected isolates in enhancing their PGP activity on tomato. This study, therefore, demonstrated that the rhizosphere of the two medicinal plant species growing in Meghalaya harbored antagonistic and PGP Actinomycetota that could be used to improve tomato crop production.
5. AUTHORS’ CONTRIBUTIONS
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agreed to be accountable for all aspects of the work. All the authors are eligible to be an author as per the International Committee of Medical Journal Editors (ICMJE) requirements/guidelines.
6. FUNDING
This work was partially funded by the Department of Biotechnology, Government of India, New Delhi under the DBT-STAR College Scheme for the year 2017-2018 for the promotion of students’ research in colleges.
7. CONFLICTS OF INTEREST
The authors report no financial or any other conflicts of interest in this work.
8. ETHICAL APPROVALS
Since the samples used during the study were not collected from any private, protected, or culturally significant location, and since this work also did not involve any human or animal subjects, no ethical approvals were required.
9. DATA AVAILABILITY
All data generated and analyzed are included within this research article.
10. PUBLISHER’S NOTE
This journal remains neutral with regard to jurisdictional claims in published institutional affiliation.
REFERENCES
1. Munaweera TI, Jayawardana NU, Rajaratnam R, Dissanayake N. Modern plant biotechnology as a strategy in addressing climate change and attaining food security. Agric Food Secur 2022;11:26. [CrossRef]
2. Saeed Q, Xiukang W, Haider FU, Kucerik J, Mumtaz MZ, Holatko J,
3. Matilla MA, Krell T. In:Egamberdieva D, Ahmad P, editors. Plant growth promotion and biocontrol mediated by plant-associated bacteria. In:Plant Microbiome:Stress Response. Microorganisms for Sustainability. Vol. 5. Singapore:Springer;2018. 45-80. [CrossRef]
4. Oren A, Garrity GM. Valid publication of the names of forty-two phyla of prokaryotes. Int J Syst Evol Microbiol 2021;71:005056. [CrossRef]
5. Goodfellow M. In:Goodfellow M, Kämpfer P, Busse HJ, Trujillo ME, Suzuki KI, Ludwig W,
6. Genilloud O. Actinomycetes:Still a source of novel antibiotics. Nat Prod Rep 2017;34:1203-32. [CrossRef]
7. Wang Z, Solanki MK, Yu ZX, Anas M, Dong DF, Xing YX,
8. Trinidad-Cruz JR, Rincón-Enríquez G, Evangelista-Martínez Z, Guízar-González C, Enríquez-Vara JN, López-Pérez L,
9. Laassami A, Yekkour A, Meklat A, Djemouai N, Zitouni A, Mokrane S,
10. Roy S, Santra HK, Banerjee D. In:Yadav AN, Rastegari AA, Yadav N, Kour D, editors. Diversity and biotechnological potential of culturable rhizospheric actinomicrobiota. In:Advances in Plant Microbiome and Sustainable Agriculture:Microorganisms for Sustainability. Vol. 19. Singapore:Springer Nature;2020. 159-87. [CrossRef]
11. Anusree T, Bhai RS, Shabeer TP, Oulkar D.
12. Köberl M, Schmidt R, Ramadan EM, Bauer R, Berg G. The microbiome of medicinal plants:Diversity and importance for plant growth, quality, and health. Front Microbiol 2013;4:400. [CrossRef]
13. Djemouai N, Meklat A, Gaceb-Terrak R, Youcef KO, Nacer A, Saadi SA,
14. Janatiningrum I, Lestari Y. Enzyme production, antibacterial and antifungal activities of
15. Devi S, Sharma P, Rana A, Pal J, Kumari A. Diversity and plant growth-promoting potential of actinomycetes associated with the rhizosphere of
16. Khaniya L, Bhattarai R, Jan HA, Hussain W, Abbasi AM, Bussmann RW,
17. Sengupta R, Dash SS. Invasion status of three non-native species from family
18. Wahyuono S, Vairappan CS, Mohd. DM, Fauziati V, Purwantiningsih, Marchaban. Major metabolites of methylripariochromene-A, a bioactive substance in the leaves of
19. Walkley A, Black IA. An examination of the degtjareff method for determining soil organic matter, and a proposed modification of the chromic acid titration method. Soil Sci 1934;37:29-38. [CrossRef]
20. Cross T. In:Holt JG, Sneath PH, Krieg NR, editors. Growth and Examination of Actinomycetes:Some Guidelines. Bergey's Manual of Determinative Bacteriology. 9th ed. Baltimore:Williams and Wilkins;1994. 605-9.
21. Chen X, Jiang Y, Li Q, Han L, Jiang C. In:Dhanasekaran D, Jiang Y, editors. Molecular phylogenetic identification of
22. Monciardini P, Sosio M, Cavaletti L, Chiocchini C, Stefano D. New PCR primers for the selective amplication of 16S rDNA from different groups of actinomycetes. FEMS Microbiol Ecol 2002;42:419-29. [CrossRef]
23. Farris MH, Olson JB. Detection of actinobacteria cultivated from environmental samples reveals bias in universal primers. Lett Appl Microbiol 2007;45:376-81. [CrossRef]
24. Mingma R, Pathom-Aree W, Trakulnaleamsai S, Thamchaipenet A, Duangmal K. Isolation of rhizospheric and roots endophytic actinomycetes from Leguminosae plant and their activities to inhibit soybean pathogen,
25. Syiemiong D, Jha DK. Antibacterial potential of
26. Vijayakumar R, Vaijayanthi G, Panneerselvam A, Thajuddin N. In:Dhanasekaran D, Thajuddin N, Panneerselvam A, editors.
27. Khamna S, Yokota A, Lumyong S. Actinomycetes isolated from medicinal plant rhizosphere soils:Diversity and screening of antifungal compounds, indole-3-acetic acid and siderophore production. World J Microbiol Biotechnol 2009;25:649-55. [CrossRef]
28. Syiemiong D, Jha DK. Search for plant growth promoting actinobacteria from a limestone mining spoil soil in Meghalaya. Res J Life Sci Bioinformatics Pharm Chem Sci 2019;5:1024-36.
29. Tamreihao K, Nimaichand S, Chanu SB, Devi KA, Lynda R, Jeeniita N,
30. Jog R, Nareshkumar G, Rajkumar S. Plant growth promoting potential and soil enzyme production of the most abundant
31. Thampi A, Bhai RS. Rhizosphere actinobacteria for combating
32. Momin MD. Characterization of Rhizospheric Actinomycetes of Major Crop Plants and their Plant Growth Promoting Properties Under Jhum fields of Mizoram. Aizawl:Mizoram University;2020.
33. Ostash B, Gren T, Hrubskyy Y, Tistechok S, Beshley S, Baranov V,
34. Palaniyandi SA, Yang SH, Damodharan K, Suh JW. Genetic and functional characterization of culturable plant-beneficial
35. Maciejewska M, Adam D, Martinet L, NaôméA, Ca?usinska M, Delfosse P,
36. Tan HM, Cao LX, He ZF, Su GJ, Lin B, Zhou SN. Isolation of endophytic actinomycetes from different cultivars of tomato and their activities against
37. Alekhya G, Gopalakrishnan S. Exploiting plant growth-promoting
38. Thawai C.
39. Ningthoujam DS, Lynda RK, Tamreihao K, Chanu SB, Aruna KH, Jeeniita N. Isolation and characterization of
40. De Brito AM, Gagne S, Antoun H. Effect of compost on rhizosphere microflora of the tomato and on the incidence of plant growth-promoting rhizobacteria. Appl Environ Microbiol 1995;61:194-9. [CrossRef]
41. Guo X, Liu N, Li X, Ding Y, Shang F, Gao Y,
42. Sharma P, Thakur D. Antimicrobial biosynthetic potential and diversity of culturable soil actinobacteria from forest ecosystems of Northeast India. Sci Rep 2020;10:4104. [CrossRef]
43. Nimaichand S, Devi AM, Tamreihao K, Ningthoujam DS, Li WJ. Actinobacterial diversity in limestone deposit sites in Hundung, Manipur (India) and their antimicrobial activities. Front Microbiol 2015;6:413. [CrossRef]
44. Bauermeister A, Calil FA, Pinto FD, Medeiros TC, Almeida LC, Silva LJ,
45. Hashizume H, Sawa R, Yamashita K, Nishimura Y, Igarashi M. Structure and antibacterial activities of new cyclic peptide antibiotics, pargamicins B, C and D, from
46. Izuta S, Kosaka S, Kawai M, Miyano R, Matsuo H, Matsumoto A,
47. Msimbira LA, Smith DL. The roles of plant growth promoting microbes in enhancing plant tolerance to acidity and alkalinity stresses. Front Sustain Food Syst 2020;4:106. [CrossRef]
48. De la Torre-Hernández ME, Salinas-Virgen LI, Aguirre-Garrido JF, Fernández-González AJ, Martínez-Abarca F, Montiel-Lugo D,
49. He A, Niu S, Yang D, Ren W, Zhao L, Sun Y,
50. Tkacz A, Cheema J, Chandra G, Grant A, Poole PS. Stability and succession of the rhizosphere microbiota depends upon plant type and soil composition. ISME J 2015;9:2349-59. [CrossRef]
51. Al-Dhabi NA, Esmail GA, Ghilan AK, Arasu MV. Composting of vegetable waste using microbial consortium and biocontrol efficacy of
52. Barman D, Dkhar MS. Plant growth-promoting potential of endophytic bacteria isolated from
53. Barman D, Dkhar MS. Seasonal variation influence endophytic actinobacterial communities of medicinal plants from tropical deciduous forest of Meghalaya and characterization of their plant growth-promoting potentials. Curr Microbiol 2020;77:1689-98. [CrossRef]
54. Passari AK, Upadhyaya K, Singh G, Abdel-Azeem AM, Thankappan S, Uthandi S,
55. Dochhil H, Dkhar MS, Barman D. Seed germination enhancing activity of endophytic
56. Hassani MA, Durán P, Hacquard S. Microbial interactions within the plant holobiont. Microbiome 2018;6:58. [CrossRef]
57. Arif MS, Shahzad SM, Riaz M, Yasmeen T, Shahzad T, Akhtar MJ,